Automated identification of pattani local medicinal herbs based on deep learning techniques
Main Article Content
Abstract
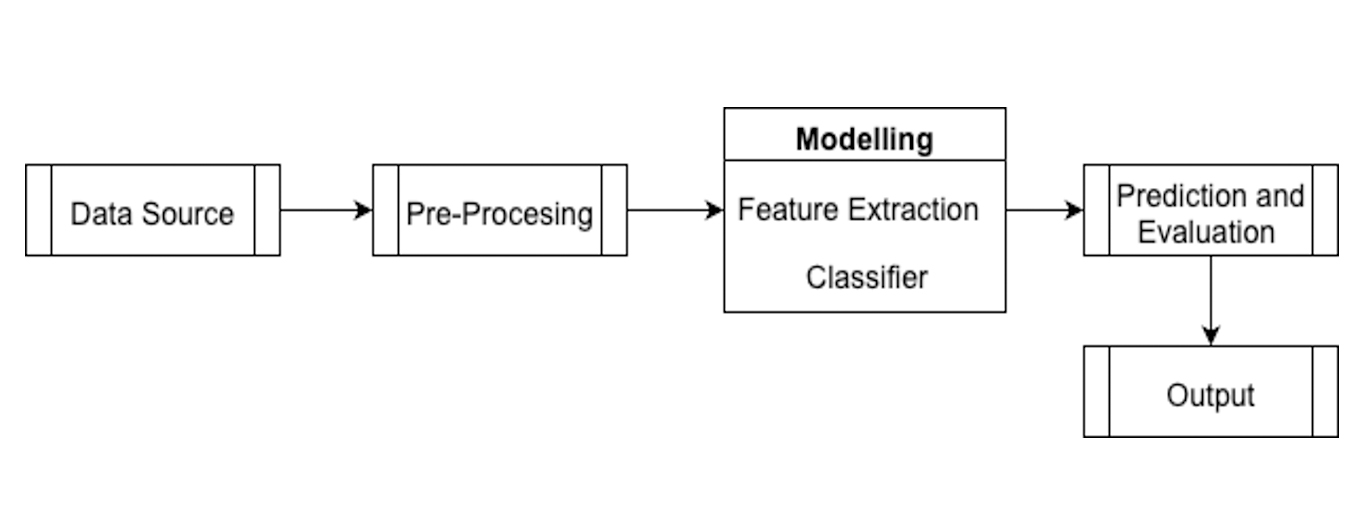
Despite Pattani's rich biodiversity, there is a significant lack of digital datasets and automated tools to preserve local ethnomedicinal knowledge. Traditionally, herbal medicine plays a significant role in healthcare systems, especially in regions with rich biodiversity, such as Pattani, Thailand. However, the manual identification of medicinal herbs is often labor-intensive and disposed to inaccuracies. This study introduces an automated system for identifying Pattani's local medicinal herbs (Etlingera elatior, Euphorbia hirta, and Leucas aspera) using Deep Learning methods, specifically Convolutional Neural Networks (CNNs). The system uses image processing techniques to improve the accuracy and confidence in herb identification. The system uses a Convolutional Neural Network (CNN) architecture comprising feature-extraction layers with ReLU activation and max pooling, followed by a fully connected softmax classifier. Data augmentation techniques were employed to enhance model generalization on the collected dataset. It additionally protects traditional knowledge through scientific validation. Collecting and pre-processing a dataset of 600 images and using CNNs yielded an overall test accuracy of 97%. Such performance reinforces the system's capabilities in healthcare for traditional practitioners and pharmaceutical researchers who need precise herb identification. Integrating technology and local knowledge to use and conserve Pattani's medicinal plants and to harness local medicinal plant knowledge is a crucial step this research takes.
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
References
Abera B. Medicinal plants used in traditional medicine by Oromo people, Ghimbi District, Southwest Ethiopia. J Ethnobiol Ethnomed. 2014;10(1):1-15.
Kumar A, Ramchiary N, Singh PJS. Role of traditional ethnobotanical knowledge and indigenous communities in achieving sustainable development goals. Sustainability. 2021;13(6):3062.
Mulugeta AK, Sharma DP, Mesfin AH. Deep learning for medicinal plant species classification and recognition: A systematic review. Front Plant Sci. 2024;14:1286088.
Chen SL, Yu H, Luo HM, Wu Q, Li CF, Steinmetz A. Conservation and sustainable use of medicinal plants: problems, progress, and prospects. Chin Med. 2016;11:37.
Bhujun V, Awan AT, Baran Y, Bunnefeld N, Chan K, Dela Cruz TE, et al. Biodiversity, drug discovery, and the future of global health: Introducing the biodiversity to biomedicine consortium, a call to action. J Glob Health. 2017;7(2):020304.
Tilburt JC, Kaptchuk TJ. Herbal medicine research and global health: An ethical analysis. Bull World Health Organ. 2008.
Sulaiman MR, Mazlan R. The ethnobotanical importance of Etlingera elatior in Southeast Asian traditional medicine. Borneo J Resour Sci Technol. 2020;3(2):101-9.
Rahmat A, Ahmad M, Salleh N. Medicinal uses of Etlingera elatior in Malaysian traditional medicine. J Ethnopharmacol. 2019;25(2):185-94.
Ghasemzadeh A, Jaafar HZ, Rahmat A. Antioxidant activities, phytochemicals, and bioactive compounds of torch ginger (Etlingera elatior). J Food Sci. 2015;80(5):956-65.
Zuraidah, Widiasari, Oviana W. Antibacterial potential of leaf and fruit of citrus extract (Citrus aurantifolia) against Klebsiella oxytoca. JPBIO J Pendidik Biol [internet]. 2023;8(1):1-11. Available from: http://jurnal.stkippersada.ac.id/jurnal/index.php/JBIO/index
Azlina AB, Noor AN, Azlan SA. Culinary uses of Etlingera elatior (Torch ginger) in Southeast Asia. J Ethnobiol Ethnomed. 2018;14(1):45.
Bhowmik D, Kumar KPS, Chiranjib, Tiwari P. Traditional and medicinal uses of Euphorbia hirta. J Chem Pharm Res. 2013;5(2):117-21.
Gohil P, Patel M, Sharma P. Euphorbia hirta: A review on its phytochemical, ethnomedicinal, and pharmacological profile. Pharmacol Rev. 2010;4(3):58-61.
Rizvi SMD, Mishra R. Gastroprotective effect of Euphorbia hirta: Ethnobotanical and pharmacological relevance. Asian Pac J Trop Dis. 2014;4(1):255-61.
Lohani M, Joshi D. Euphorbia hirta latex: Medicinal applications and phytochemistry. J Herb Med. 2020;24:100386.
Prajapati MS, Patel JB, Modi K, Shah MB. Leucas aspera: A review. Pharmacogn Rev. 2010;4(7):85-7. doi: 10.4103/0973-7847.65330.
Chandrasekaran M, Venkatesalu V. Antibacterial and anti-fungal activity of Leucas aspera leaf extracts. Asian Pac J Trop Biomed. 2010;2(4):246-9.
Gandhisan R, Thamaraichelvan A, Baburaj S. Anti-inflammatory action of Leucas aspera in rats and mice. Indian J Pharmacol. 2012;24(1):75-8.
Murti K, Kumar U. Leucas aspera: A review on its botany, phytochemical, and pharmacological profile. Int J Pharm Pharm Sci. 2012;4(3):47-51.
Pholdee N, Tokaew W. Local wisdom of medicinal plants for health care. Maha Sarakham: Rajabhat Maha Sarakham University; 2017.
Zaheer R, Shaziya H. A study of the optimization algorithms in deep learning. In: Proc 3rd Int Conf Inventive Syst Control (ICISC). 2019. p. 536-9. doi:10.1109/ICISC44355.2019.9036442.
LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521(7553):436-44.
Xie L, Wang J, Wei Z, Tian Q. Genetic CNN. Johns Hopkins Univ; 2017.
Lu J, Tan L, Jiang H. Review on convolutional neural network (CNN) applied to plant leaf disease classification. Xihua Univ. 2021.
Zin IAM, Ibrahim Z, Dino I, Aliman S, Sabri N, Mangshor NA. Herbal plant recognition using deep convolutional neural network. Universiti Teknologi MARA Malaysia; 2020.
Narkhede S. Understanding confusion matrix [internet]. Towards Data Science. 2018. Available from: https://towardsdatascience.com/understanding-confusion-matrix-a9ad42dcfd62.
Kumar A, Sharma G, Sharma A, Chopra P, Rattan P, editors. Advances in Networks, Intelligence and Computing: Proceedings of the International Conference on Networks, Intelligence and Computing (ICONIC 2023). 1st ed. Boca Raton: CRC Press; 2024.
Deng L, Yu D. Deep learning methods for feature extraction in neural networks. J Signal Process Syst. 2023;92(1):103-20.
Zhang L, Wang H, Ma Y. The role of pooling layers in CNNs: A comprehensive review. J Comput Vis Appl. 2023;34(5):432-45.
Srivastava N, Hinton G, Krizhevsky A, Salakhutdinov R. Dropout: A method to prevent neural networks from overfitting. J Mach Learn Res. 2023;16(1):1929-58.
Abadi M, Barham P, Chen J, Chen Z, Davis A, Dean J, et al. TensorFlow: A system for large-scale machine learning [internet]. In: Proc 12th USENIX Symp Operating Syst Design Implementation (OSDI' 16). 2016. p. 265-83. Available from: https://www.usenix.org/conference/osdi16/technical-sessions/presentation/abadi.
Wang F, Jiang MQ, Qian C, Yang S, Li C, Zhang HG, et al. Residual attention network for image classification. In: Proceedings of 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2017. p. 6450-6458.
Chollet F. Deep learning with Python. New York: Manning Publications; 2017.
Goodfellow I, Bengio Y, Courville A. Deep learning. Cambridge: MIT Press; 2016.
Gu X, Yang X, Li J. Enhancing feature extraction in convolutional neural networks. IEEE Trans Image Process. 2023;32(2):156-68.
Thanapatpisarn P. Fundamental Data Analytics & Data Scientist EP.20 [internet]. Medium. 2021 Nov 27. Available from: https://datascihaeng.medium.com/evaluation-matrix-part1-ad629e648f8
Gajendran MK, Rohowetz LJ, Koulen P, Mehdizadeh A. Novel machine-learning-based framework using electroretinography data for the detection of early-stage glaucoma. Front Neurosci. 2022;16:869137.