Drug Selection Support System for Targeted Cancer Therapy Based on Cell Signal Transduction Pathways
Main Article Content
Abstract
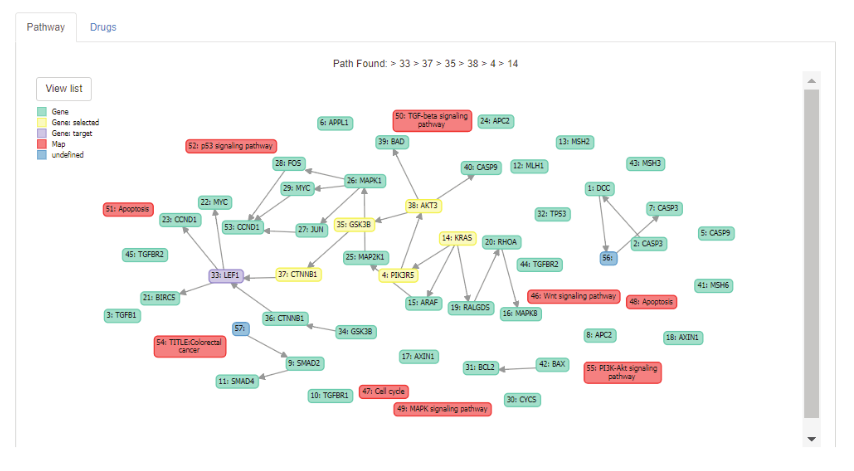
This research proposed drug selection support system for targeted cancer therapy based on cell signal transduction pathways. Two graph traversal algorithms include depth first search (DFS) and breadth first search (BFS) algorithms were brought to apply with centrality measurements. There are called centrality depth first search (CDFS) algorithm. The CDFS was used to traverse in the cell signal transduction pathway for genes that related with the interested gene in the same pathway. Then the system will bring that genes to find drugs that can be used for targeted cancer therapy. Moreover, this research also developed scoring drug system for computing score of that drugs and ranking drugs according to their score. The results shown that drug selection support system for targeted cancer therapy based on signal transduction pathway can decrease time to analyze pathway and find drugs based on cell signal transduction pathway.
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
All authors need to complete copyright transfer to Journal of Applied Informatics and Technology prior to publication. For more details click this link: https://ph01.tci-thaijo.org/index.php/jait/copyrightlicense
References
Bayat, M.R. et al. (2017). Combination therapy in combating cancer. Oncotarget. 8(23). 38022-38043.
Bidgoli, H. (1989). Decision Support Systems: Principles and Practices. St. Paul: West Publishing Company. 1989.
BNH. (2011). Cancer Treatment Trends. Better Health Magazine Issue 2. Available from : https://www.bumrungrad.com/en/betterhealth/2011/cancer-treatments/cancer-treatment-trends. [August 18, 2011].
BNH. (2013). Targeted Therapy. Health Spot. Available from: https://www.bumrungrad.com/healthspot/November-2013. [August 36, 2013].
ClinicalTrials.gov. (2015). Available from: https://clinicaltrials.gov. [February 26, 2015].
Petrochilos, D. and Abernethy, N. (2012). Assessing Network Characteristics of Cancer Associated Genes in Metabolic and Signaling Networks. Proceeding of the 2012 IEEE Symposium on Computational Intelligence in Bioinformatics and Computational Biology (CIBCB), San Diego, CA, USA, May 9-12, 2012, 290-297.
FDA@Drugs (2015). Available from: https://www.accessdata.fda.gov. [February 26, 2015].
Keen, P.G.W. and Scott Morton, M.S. (1978). Decision Support Systems: An Organization Perspective. Reading. MA : Addison-Wesley. 1978.
KEGG. (2015). KEGG CANCER. Available from: https://www.genome.jp/kegg. [February 26, 2015].
Dayeh, M. and Hahsler, M. (2012). Biological Pathway Completion using Network Motifs and Random Walks on Graphs. Proceeding of the 2012 IEEE Symposium on Computational Intelligence in Bioinformatics and Computational Biology (CIBCB), San Diego, CA, USA, May 9-12, 2012, 229-236.
Tsai, M. et al. (2010). Genetic Regulatory Networks Established by Shortest Path Algorithm and Conditional Probabilistic for Ovarian Carcinoma Microarray Data. Proceeding of the 9th International Conference on Machine Learning and Cybernetics, Qingdao, July 11-14, 2010, 32-36.
National Cancer Institute (2017). NCI Dictionary of Cancer Terms. Available from: https://www.cancer.gov/publications/dictionaries/cancer-terms. [January 10, 2018].
National Cancer Institute (2017). Types of Cancer Treatment. Available from: https://www.cancer.gov/about-cancer/treatment/types. [January 10, 2018].
Pankaj Agarwal and David B. Searls. (2008). Literature Mining in Support of Drug Discovery. Briefings in Bioinformatics. 9(6), 479-492.
Taysir Hassan A. Soliman et al. (2012). Mining Disease Integrated Ontology. Proceeding of the 2012 IEEE 12 th International Conference on Bioinformatics and Bioengineering (BIBE), Larnaca, Cyprus, November 11-13, 2012, 40-45.
Turban, E et al. (2005). Introduction to Information Technology. Hoboken : John Wiley & Sons. 2005.
WHO. (2015). World health report. Available from: https://www.who.int. [January 12, 2015].
Yang, y. et al. (2009). Target Discovery from Data Mining Approaches. Drug Discovery Today. 14, 147-154.