Comparison of CNN Architectures for Thai Medicinal Plant Classification
Main Article Content
Abstract
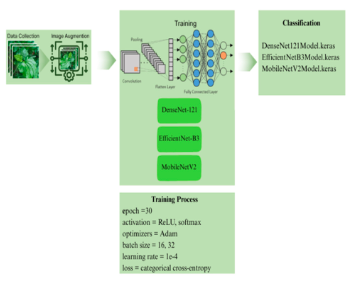
Thai medicinal plants are essential to traditional healthcare and local livelihoods. However, many Thai medicinal plants have similar morphological characteristics such as shape, colour, and texture. This problem leads to misidentification and misclassification. Image classifiers utilizing convolutional neural networks (CNNs), which are a class of deep learning models, provide a scalable substitute for manual classification. This study aims to evaluate and compare the performance of three CNN architectures (DenseNet-121, EfficientNet-B3, and MobileNetV2) for classifying 10 species of Thai medicinal plants. The dataset comprises 5,000 leaf images representing 10 species (500 images per species). This study partitioned the dataset into 80% training set and a 20% test set. To enhance model generalization, we applied data augmentation techniques-specifically rotation, flipping, and colour manipulation. Furthermore, we utilized TensorFlow and Keras on Google Colab with GPU acceleration to train the models. Evaluation metrics include accuracy, precision, recall, F1 score, model size, inference time, and CPU utilization. The results highlight a trade-off between accuracy and efficiency: DenseNet-121 achieved the highest accuracy at 96.0% and a Matthews Correlation Coefficient (MCC) of 0.9558. Statistical analysis confirmed that DenseNet-121 significantly outperformed the other architectures (p < 0.05), albeit with a higher inference time (579.22 s). Notably, EfficientNet-B3 and MobileNetV2 both achieved an accuracy of 93.4%, with MobileNetV2 performing the best in terms of model size (11.07 MB) and inference time (3.86 s). In conclusion, DenseNet-121 is the most accurate model, while MobileNetV2 is best suited for real-time applications due to its lightweight and rapid inference time. EfficientNet-B3 offers an optimal balance between accuracy and computational efficiency.
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
References
W. Ruangaram and E. Kato, “Selection of Thai medicinal plants with anti-obesogenic potential via in vitro methods,” Pharmaceuticals, vol. 13, no. 4, p. 56, 2020.
R. Geerthana, P. Nandhini and R. Suriyakala, “Medicinal plant identification using deep learning,” International Research Journal of Advanced Science Hub, vol. 3, no. 05S, pp. 48–53, 2021.
N. Duong-Trung, L. D. Quach, M. H. Nguyen and C. N. Nguyen, “A combination of transfer learning and deep learning for medicinal plant classification,” in Proceedings of the 4th International Conference on Intelligent Information Technology, pp. 83–90, 2019.
R. Azadnia, M. M. Al-Amidi, H. Mohammadi, M. A. Cifci, A. Daryab and E. Cavallo, “An AI based approach for medicinal plant identification using deep CNN based on global average pooling,” Agronomy, vol. 12, no. 11, p. 2723, 2022.
R. Azadnia, F. Noei-Khodabadi, A. Moloudzadeh, A. Jahanbakhshi and M. Omid, “Medicinal and poisonous plants classification from visual characteristics of leaves using computer vision and deep neural networks,” Ecological Informatics, vol. 82, p. 102683, 2024.
S. K. J. Urumarudappa et al., “Development of a DNA barcode library of plants in the Thai Herbal Pharmacopoeia and monographs for authentication of herbal products,” Scientific Reports, vol. 12, no. 1, 2022.
S. S. Keh, “Semi-supervised Noisy Student pre-training on EfficientNet architectures for plant pathology classification,” arXiv preprint, arXiv:2012.00332, 2020.
K. Labrighli, C. Moujahdi, J.E. Oualidi and L. Rhazi, “Artificial intelligence for automated plant species identification: a review,” International Journal of Advanced Computer Science and Applications, vol. 13, no. 10, 2022.
Y. LeCun, Y. Bengio and G. Hinton, “Deep learning,” Nature, vol. 521, no. 7553, pp. 436–444, 2015.
A. K. Mulugeta, D. P. Sharma and A. H. Mesfin, “Deep learning for medicinal plant species classification and recognition: a systematic review,” Frontiers in Plant Science, vol. 14, p. 1286088, 2024.
W. Huang and H. Zhang, “Convergence analysis of deep residual networks,” Analysis and Applications, vol.22, no.2, pp. 351-382, 2023.
A. Dosovitskiy et al., “An Image is Worth 16×16 Words: Transformers for Image Recognition at Scale,” arXiv preprint, arXiv:2010.11929, 2021.
G. Huang, Z. Liu, L. Van Der Maaten and K. Q. Weinberger, “Densely Connected Convolutional Networks,” 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, pp. 2261-2269, 2017.
M. Tan and Q. Le, “EfficientNet: rethinking model scaling for convolutional neural networks,” IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 42, no. 8, pp. 6105–6114, 2019.
M. Sandler, A. Howard, M. Zhu, A. Zhmoginov and L. -C. Chen, “MobileNetV2: Inverted Residuals and Linear Bottlenecks,” 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, pp. 45104520, 2018.
N. A. M. Roslan, N. M. Diah, Z. Ibrahim, Y. Munarko and A. E. Minarno, “Automatic plant recognition using convolutional neural network on Malaysian medicinal herbs: the value of data augmentation,” International Journal of Advanced Intelligent Informatics, vol. 9, no. 1, pp. 136–144, 2023.
M. T. Ahad, Y. Li, B. Song and T. Bhuiyan, “Comparison of CNN-based deep learning architectures for rice diseases classification,” Artificial Intelligence in Agriculture, vol. 9, pp. 22–35, 2023.
E. C. Too, L. Yujian, S. Njuki and L. Yingchun, “A comparative study of fine-tuning deep learning models for plant disease identification,” Computers and Electronics in Agriculture, vol. 161, pp. 272–279, 2019.